"We mix 10kg of passion, then add 5000 spatulas full of hard work. We heat our portion with a warmth of confidence and run the mixture in a perseverance gel. We stain for patience and resilience in a buffer full of success! We see the bands and marks of millions of accomplishments shining strong under the light of learning. We transfer our knowledge into marks of aspiration and excellence that show today and tomorrow, all the way."
A tumor is an evolving ecosystem. The dynamic interplay between heterogeneous tumor cells and their surrounding tissue microenvironment (TME) governs how tumors initiate, progress, and ultimately spread to distant organs. Our lab investigates the co-evolution between tumor cells and the TME during therapeutic resistance and metastatic relapse, integrating cancer biology, functional genomics, single-cell technologies, and computational approaches to develop precision strategies against cancer.
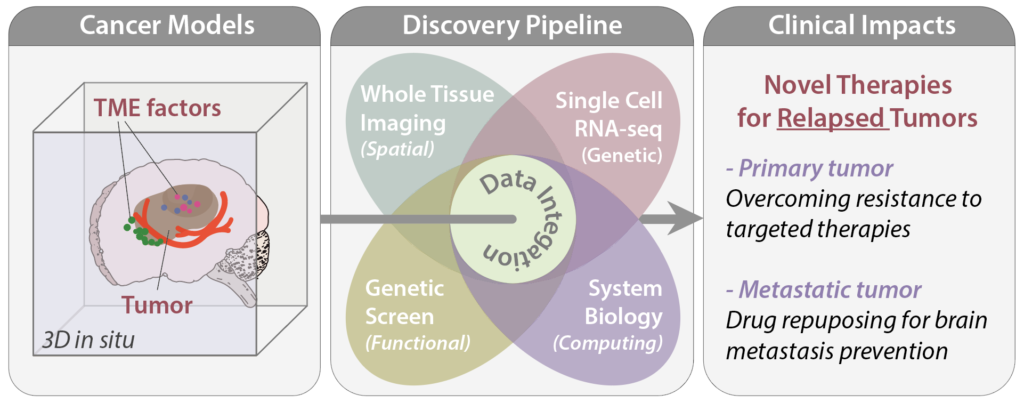
Theme 1: Tumor Evolution and Microenvironment Co-evolution
Resistance to targeted therapy represents one of the most significant clinical challenges in oncology. Rather than arising from isolated mutations, resistance emerges from continuous evolutionary adaptation to a complex, dynamic TME. Our lab studies this process by mapping tumor evolution trajectories in transgenic mouse models, examining how selection pressures such as chemotherapy and targeted therapy reshape the tumor immune landscape and influence metastasis and immunotherapy response (Wang et al., Nature Communications 2019).
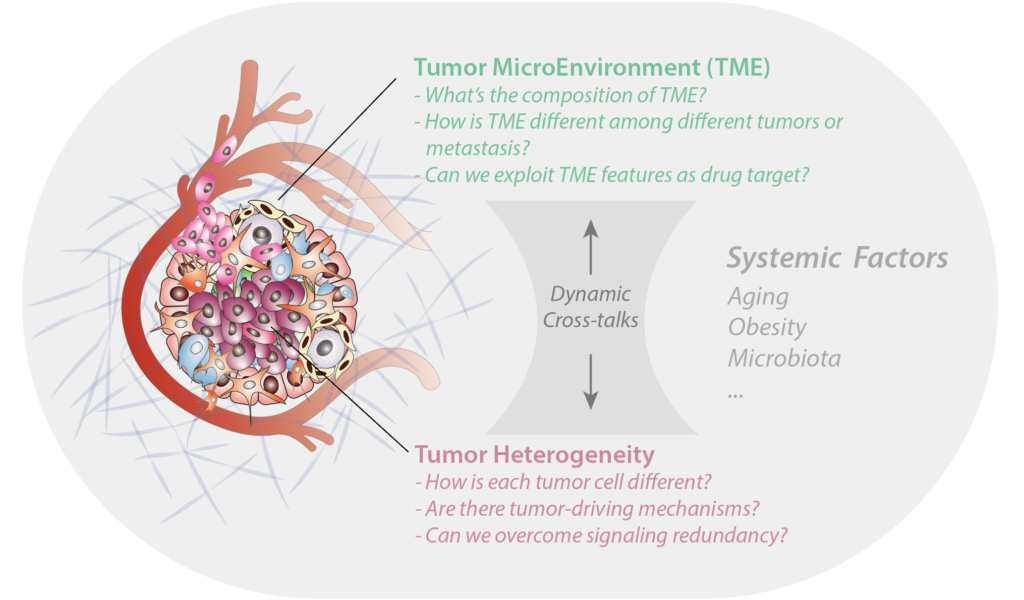
A central challenge is that the co-evolution between tumor and TME is not governed by a handful of discrete genetic or epigenetic events, but by highly plastic, continuous transcriptome trajectories shaped by redundant, non-linear regulatory circuits. We ask whether, within this complexity, there are identifiable and tractable trajectories that govern tumor immunity. To address this, we apply multiplexed genetic perturbation approaches, including Perturb-seq and CROP-seq, and combine them with machine learning-based systems biology to investigate signaling plasticity and redundancy during tumor-TME interactions. Using single-cell RNA sequencing (CITE-seq) and spectral flow cytometry, we resolve the enormous cellular heterogeneity of the TME and generate testable hypotheses about tumor immunity that inform novel therapeutic strategies (Palakurthi et al., Nature Communications 2023).
Theme 2: Metastasis and the Niche
Cancer metastasis requires tumor cells to colonize distant and drastically different tissue environments. A fundamental question is how continuously disseminating tumor cells (DTCs) survive, home to distant organs, and adapt to specialized metastatic niches, and how the reciprocal interplay between intrinsic tumor cell programs and niche-driven signals shapes this process. We investigate these questions through complementary experimental approaches spanning metabolic biology, clonal dynamics, and tumor-immune interactions.
A key finding from our lab is that early metastatic colonization involves a metabolic transcriptome shift driven by epigenetic upregulation of glutamate decarboxylase 1 (GAD1), which facilitates colonization through the GABA metabolic pathway. Repurposing the FDA-approved anti-seizure drug vigabatrin to inhibit this axis significantly reduced metastatic burden in vivo (Schnepp et al., Cancer Research 2017). To complement targeted mechanistic studies, we developed a hybrid screening strategy combining RNA profiling of temporal transcriptome changes with a rapid forward genetic screen in Drosophila to identify drivers of metastatic adaptation in an unbiased manner (Nature Communications 2020).
More recently, we developed MetTag, a single-cell barcoding and transcriptome profiling approach with time-stamped clonal identifiers, to resolve the temporal dynamics of DTC dissemination across evolving metastatic niches. In studies of ovarian cancer metastasis (bioRxiv), deep barcode sequencing revealed that early-disseminated clones are preferentially enriched across metastatic sites, establishing a “first come, first served” principle of pro-metastatic adaptation. Single-cell RNA sequencing coupled with RNA velocity analyses in ascites and metastasis-bearing omenta uncovered a distinct interferon-gamma (IFNγ)-centric transcriptional trajectory that is selectively enriched among these early seeding clones. In vivo CRISPR/Cas9 screening of metastatic niche-specific gene signatures confirmed that members of the ascites IFN signature, including Marco, Gbp2b, and Slfn1, are functionally required for peritoneal metastasis. Knockout of IFN receptor 1 (Ifngr1) in tumor cells significantly reduced metastatic burden and extended survival, and mechanistically, we found that tumor-intrinsic IFN signaling cooperates with ascites-derived tumor-associated macrophages to protect cancer cells from anoikis by activating pro-survival NFκB/TNF signaling.
In the brain metastatic niche, we use tissue clearing and SMART3D imaging to characterize morphological and functional changes in astrocytes, microglia, and vasculature during colonization.
Research Highlight: CNS-Myeloid Activation in Brain Metastasis
Beyond structural remodeling, we investigate the contributions of resident immune cells using conditional depletion and genetic knockout models, with particular focus on how metastasis-induced neuroinflammation promotes progression (Guldner et al., Cell 2020).
- Activated CNS-Myeloid Dynamics: Identification of a distinct activated myeloid population (CD86+/CX3CR1Lo/CXCL10Hi) that drives immunosuppression in the brain.
- Immune Checkpoint Synergy: Discovery that PD-L1 and VISTA cooperate on myeloid cells to inhibit T cell infiltration and activation.
- Metastatic Niche Remodeling: Spatial analysis reveals how tumor-associated microglia and macrophages physically remodel the niche to create a pro-survival environment.
More broadly, we are interested in how systemic factors, including aging, chemotherapy exposure, and gut dysbiosis, modulate immune surveillance and shape the pre-metastatic niche. Related questions include the role of the meningeal and lymphatic systems in immune regulation of metastasis, strategies for modeling aged tissue microenvironments in vivo, and opportunities to exploit the spatial architecture of the tumor-immune interface for targeted drug delivery or imaging.
Theme 3: Epistatic Genetic Interactions and Metastatic Fitness
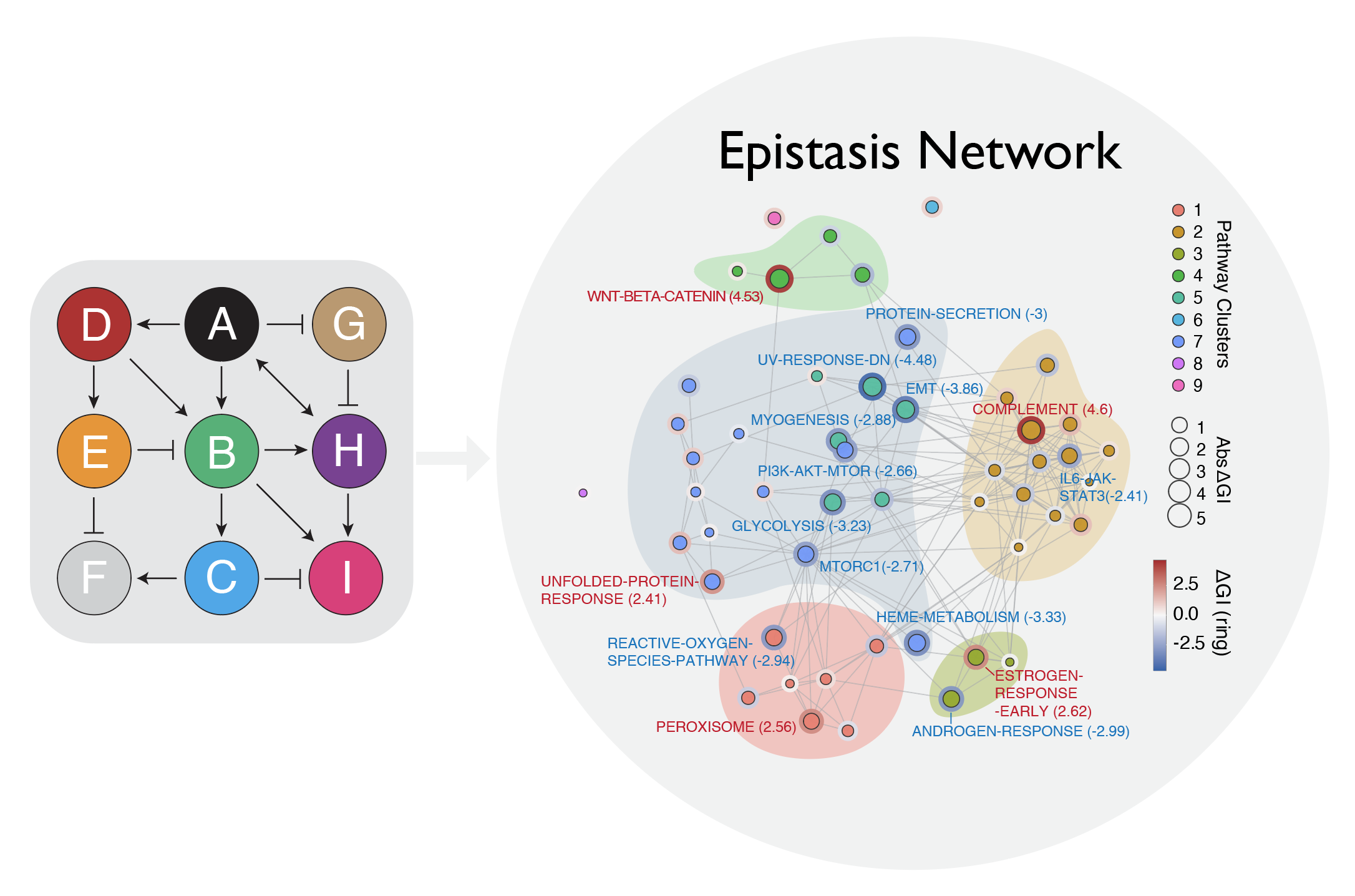
Metastatic fitness is not determined by single gene effects but by higher-order genetic networks that rewire cellular plasticity programs in response to environmental context. We have developed an innovative platform that integrates multiplexed in vivo dual-gene perturbations with single-cell transcriptomics to systematically map the epistatic networks governing metastasis. This approach captures both phenotypic outcomes and underlying transcriptional states at single-cell resolution within intact tissue, revealing how genetic interactions are context-dependent: they vary across distinct tumor subpopulations and are modulated by the local immune microenvironment.
This work demonstrates that metastatic fitness emerges from combinatorial genetic complexity that cannot be understood gene by gene. Our platform provides a generalizable framework for decoding context-dependent vulnerabilities in cancer, enabling the systematic identification of interaction nodes that shape adaptive phenotypes in vivo. By linking epistatic network structure to transcriptional state and immune context, we aim to prioritize combinatorial therapeutic targets with greater precision than single-gene approaches allow.