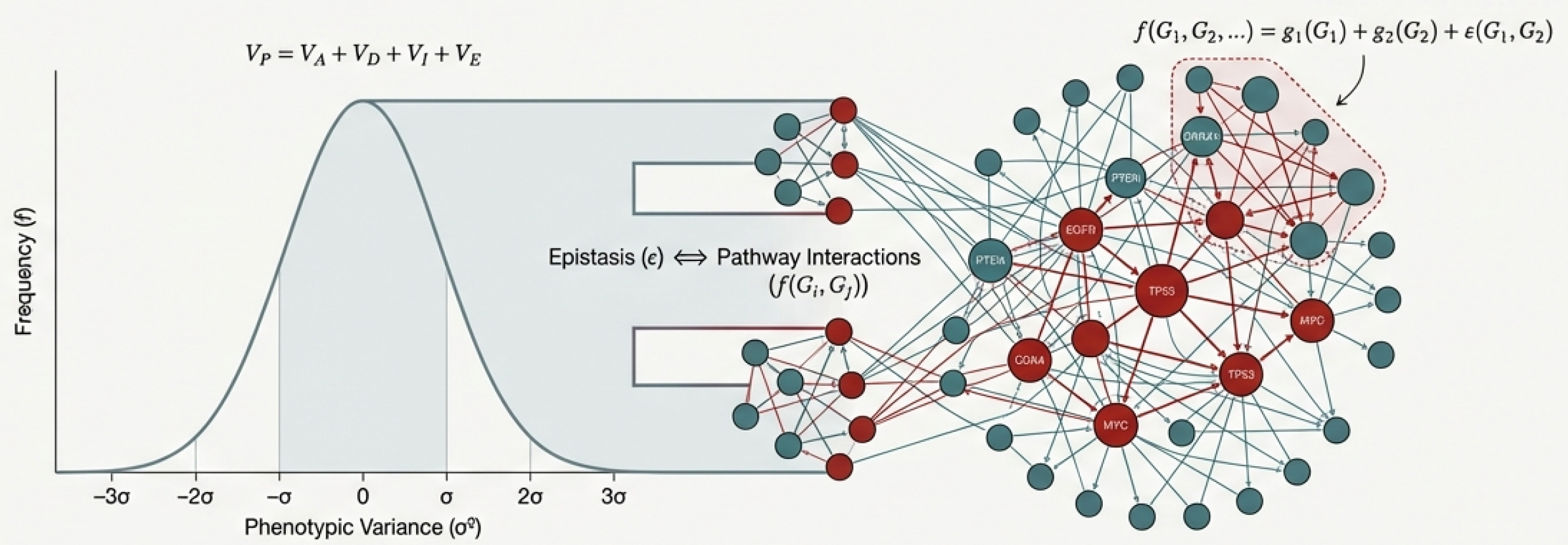
The definition of epistasis has evolved significantly since its origins, splitting into distinct frameworks depending on the field of study. Here is how population geneticists, systems biologists, and cancer biologists define and apply the concept:
Population Genetics (Statistical Epistasis)
In population genetics, epistasis is fundamentally a statistical concept. Originating from R.A. Fisher’s 1918 work, statistical epistasis is defined as the non-additive deviation in a linear model of inheritance. Rather than looking at molecular mechanisms, it measures the interaction component of genetic variance that remains after accounting for the additive effects of individual loci. It describes the interaction of allele frequencies across an entire population rather than within a single individual. Population geneticists often find this form of epistasis difficult to robustly detect in standard genome-wide association studies (GWAS) due to statistical power limitations, multiple-testing burdens, and the fact that reported signals can sometimes be artifacts of non-linear scale effects.
Systems Biology (Biological/Functional Epistasis)
Systems biologists focus on biological (or physiological) epistasis, which traces back to William Bateson’s classical definition where the effect of one gene masks or suppresses the phenotypic expression of another. Unlike the population-level statistical view, biological epistasis describes the functional and physical interaction between gene products within a single individual or cell. For systems biologists, epistasis represents the architecture of metabolic or signaling pathways. They define and map epistasis by building large-scale, global “interactomes” or genetic interaction networks using high-throughput combinatorial perturbation technologies like yeast Synthetic Genetic Arrays (SGA), CRISPR screens, and single-cell Perturb-seq.
Cancer Biology (Evolutionary and Therapeutic Epistasis)
Cancer biologists merge concepts of functional biology and evolutionary fitness landscapes to define epistasis as the governing logic of a tumor’s evolution. Instead of viewing epistasis as statistical noise, they look at how mutations interact to dictate tumor survival and vulnerabilities:
- Fitness Landscapes and Trajectories: Epistasis determines the accessible evolutionary paths a tumor can take. For example, “sign epistasis” occurs when the fitness benefit of one mutation (like KRAS) is strictly conditional on the presence of another (like STK11).
- High-Order Interactions: A tumor’s fitness is defined not just by individual driver mutations, but as an emergent property of a complex, high-order “ensemble” of three or more interacting mutations.
- Synthetic Lethality: Cancer biologists actively hunt for extreme negative epistasis, known as synthetic lethality. By understanding these network interactions, they can design rational combination therapies—such as using PARP inhibitors to exploit the epistatic vulnerability in BRCA-deficient tumors.